SURGICAL CRITICAL CARE INITIATIVE
SC2i, a consortium of federal and non-federal research institutions, develops, translates, and validates biology-driven critical care, centered on an individual patient’s biology for both short- and long-term outcomes.
We are in a unique position to leverage three things: advances in computing (large scale storage via Cloud solutions, new interfaces with EMRs), advances in machine learning (applying artificial intelligence/novel statistical techniques to derive useful information from large datasets) and explosion of biological/medical data (wide clinical and biomarker information).
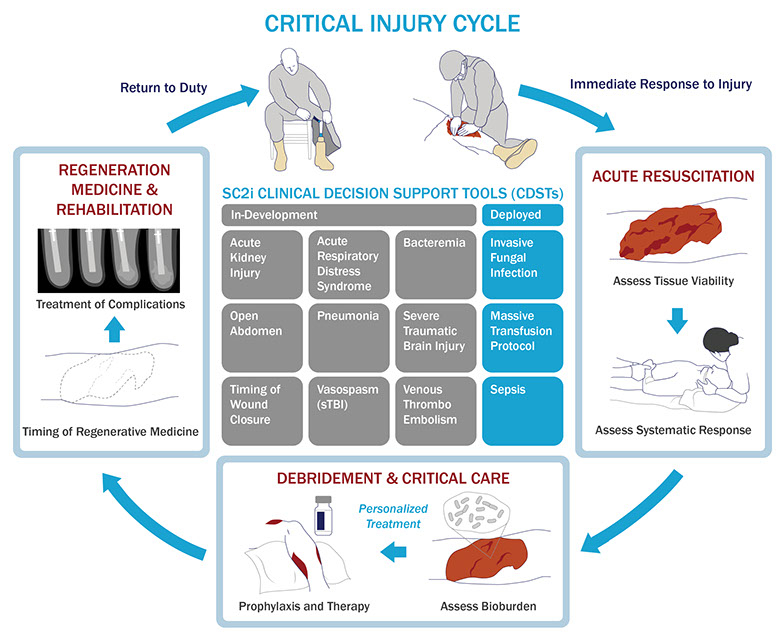
Our principal focus is to facilitate tissue acquisition and data analysis for improved decision-making algorithms. Valid results lead to the rapid integration of findings into clinical practice, thereby maximizing outcomes across any discipline requiring complex medical decision-making, including surgery, critical care, emergency medicine, orthopedics, transplant and oncology. Based on observations, and initial results from grant-funded research, we have been able to develop clinical decision support tools (CDSTs) to enhance surgical decision making in both civilian and military health systems.
Moving away from population-based risk stratification to truly personalized diagnoses and surgical interventions, to deliver the right treatment, to the right patient, at the right time - the hallmark of what we refer to as ‘precision medicine.’
OUR PROCESS
Our guiding principle, based on the Precision Medicine Initiative, is an integrative approach that focuses on individual patient biology instead of the typical population-based paradigm, which is the traditional critical care model.
We gather and analyze all available medical data for use in computerized statistical models. Clinicians and scientists at clinical facilities are invited to collaborate, using information ranging from simple observation, enrolling critically ill patients, curating biological samples, processing molecular essay, assembling clinical and biomarker data in a central data repository.
PRODUCT DEVELOPMENT
We have 40 ongoing analytics projects and three deployed CDSTs — with approximately 10 more in various stages of development—to predict the occurrence of Invasive Fungal Infection (IFI), the need for massive transfusion and Sepsis, with two of these linked to the electronic health records (EHR) at Emory University at Duke with implementation within the Military Health System.
PARTNERS
In addition to collaborating with other USU Centers, we collaborate with non-USU partners including University of Pittsburgh, University of Indiana, University of South Florida/Tampa General Hospital, Lawrence Livermore National Laboratory, Massachusetts Institute of Technology—Lincoln Laboratory, and Beth Israel Deaconess Medical Center.
These active laboratories, as well as two molecular core laboratories at USU’s Bethesda campus, and labs at the Naval Medical Research Center, Duke University, and Emory University, leverage a growing data bank of about 1,800 patients to aggressively develop nine additional predictive algorithms for conditions associated with a high-risk of morbidity and/or mortality.
The results: the development of clinical and biomarker driven Clinical Decision Support Systems (CDSS) used to improve the quality and reduce the cost of care in critically ill patients for the benefit of both military and civilian health care systems.